Michener started his talk with two contrasting points. First, we are currently deluged by data. There is more data available to scientists now than ever, perhaps 45000 exabytes by 2020. On the other hand, scientific data is constantly lost. The longer since a paper is published, the less likely its data can be recovered (one study he cited showed that data had a half life of 20 years). There are many causes of data loss, some technological, some due to changes in sharing and publishing norms. The rate at which data is lost may be declining though. We're in the middle of a paradigm shift in terms of how scientists see our data. Our vocabulary now includes concepts like 'open access', 'metadata', and 'data sharing'. Many related initiatives (e.g. GenBank, Dryad, Github, GBIF) are fairly familiar to most ecologists. Journal policies increasingly ask for data to be deposited into publicly available repositories, computer code is increasingly submitted during the review process, and many funding agencies now require statements about data management practices.
This has produced huge changes in typical research workflows over the past 25 years. But data management practices have advanced so quickly there’s a danger that some researchers will begin to feel that it is unobtainable, due to the level of time, expertise, or effort involved. I feel like sometimes data management is presented as a series of unfamiliar tools and platforms (often changing) and this can make it seem hard to opt in. It’s important to emphasize good data management is possible without particular expertise, and in the absence of cutting edge practices and tools. What I liked about Michener's talk is that it presented practices as modular ('if you do nothing else, do this') and as incremental. Further, I think the message was that this paradigm shift is really about moving from a mindset in which data management is done posthoc ('I have a bunch of data, what should I do with it?') to considering how to treat data from the beginning of the research process.
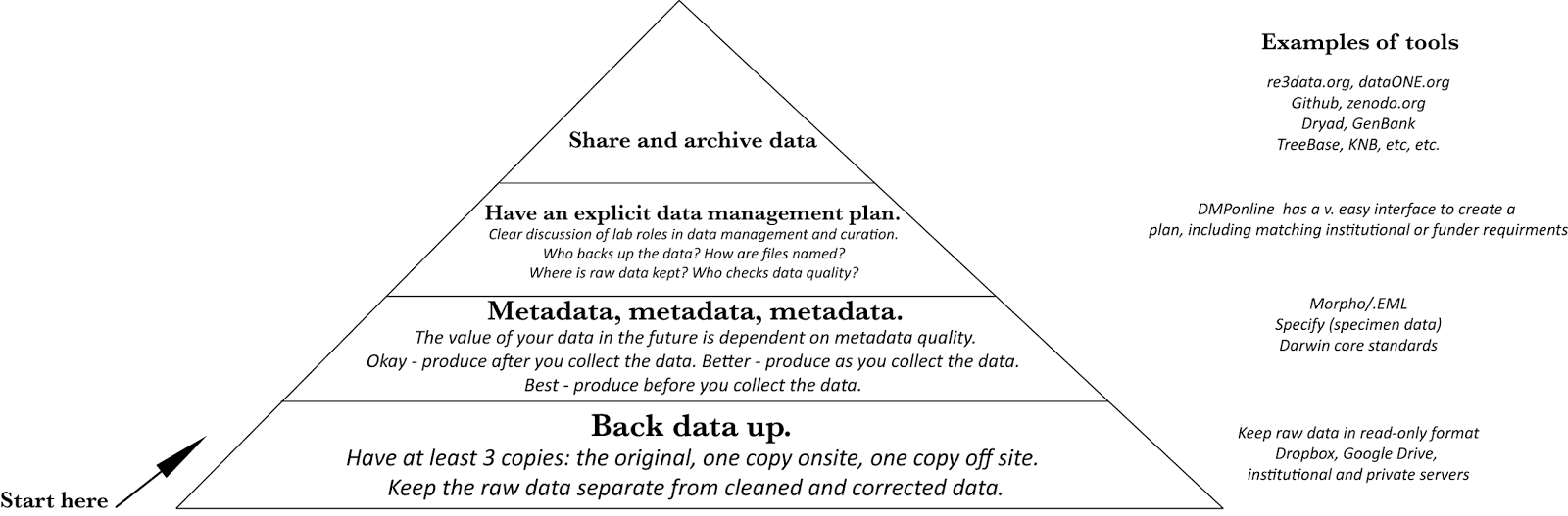 |
| Hierarchy of data management needs. |
One you make it to 'Share and archive data', you can follow some of these great references.
Hart EM, Barmby P, LeBauer D, Michonneau F, Mount S, Mulrooney P, et al. (2016) Ten Simple Rules for Digital Data Storage. PLoS Comput Biol 12(10): e1005097. doi:10.1371/journal.pcbi.1005097
James A. Mills, et al. Archiving Primary Data: Solutions for Long-Term Studies, Trends in Ecology & Evolution, Volume 30, Issue 10, October 2015, Pages 581-589, ISSN 0169-5347.
https://software-carpentry.org//blog/2016/11/reproducibility-reading-list.html (lots of references on reproducibility)
K.A.S. Mislan, Jeffrey M. Heer, Ethan P. White, Elevating The Status of Code in Ecology, Trends in Ecology & Evolution, Volume 31, Issue 1, January 2016, Pages 4-7, ISSN 0169-5347.
Thanks to Matthias Grenié for discussion on this topic.

2 comments:
THANKS! BUt still in how many universities is the concept and use of meta data from the beginning! taught? Working at a research institute with a full unite dedicated to it, researchers still hesitate to use meta data and take the effort to annotated their work properly. cheers Marten
Marten, you're exactly right. This is the exact problem we're having in our team at the cefe too. Metadata seems to be the biggest stumbling block, and I'm not sure why. A few ideas have been thrown around, including having a few templates for different types of data with set fields and just requiring every project fill the one out.
I guess if enough journals require data to be submitted it will become standard practice?...
Post a Comment